Data
Data extraction from a web source with Python + Pandas
 By Javier Andújar, in
By Javier Andújar, in Data
For the following example, we are going to use Python with Pandas in a Windows environment to do a daily load to a SQL Server database from a web source. Suppose we want to obtain information about the current COVID-19 pandemic, perhaps we want to represent the contagion risk that visiting different countries implies, so a good starting point would be to obtain some dataset that shows the number of infections discriminated by geographic regions. A good place to look for datasets is Google’s Dataset Search (https://datasetsearch.research.google.com/), the service was released in 2018 and it was out of beta as of January 23rd, 2020.
The Dataset
Searching the web, we have reached the data repository shared on GitHub by the Johns Hopkins University Center for Systems Science and Engineering, JHU CSSE COVID-19 Data, at https://github.com/CSSEGISandData/COVID-19. At the end of the page we can read that it is shared under the Creative Commons Attribution 4.0 International (CC BY 4.0) license, that is, we can use it as long as we make the corresponding attribution to the source, as described in point two of the terms of use. The source is reliable enough and they seem to have the information we need, so let us download the file and analyze it a bit.
Within all the datasets available in that repository, the one that should serve our purpose just fine is time_series_covid19_confirmed_global, a file in CSV format containing all the confirmed infections globally. The file is not large, to date it weighs 263KB. Let us open it to make a first recognition of the data and the structure:
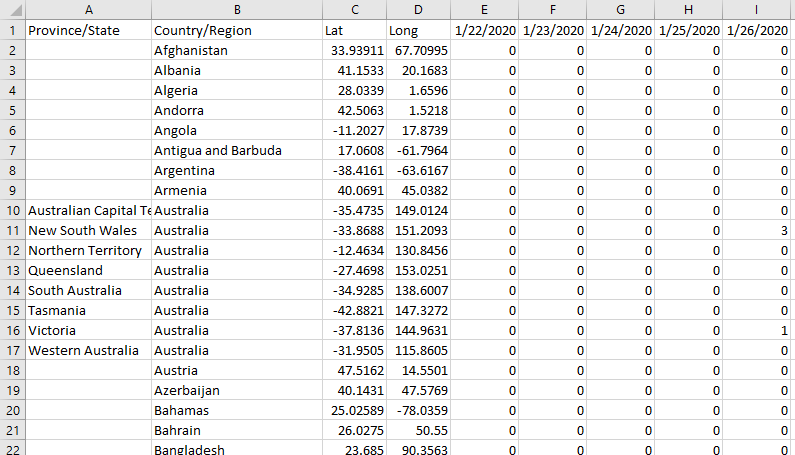
At first glance, we notice that there is a format issue: the dataset has a row for each geographical area and the number of cases is presented in successive columns, a new column is added for each day. This structure will not work for our base, because we do not want to add a new column with each load execution. Later we will have to perform some transformation to give to the data the structure we want, for that we will use Pandas. Apart from this, there is no major inconvenience with the dataset, it is very neat and there is no garbage to clean. In addition to province/state, country/region, we see that they included the latitude and longitude data, this will be important later if we want to represent the information on a map, so we are going to grab those two fields also.
Target structure
In the COVID-19 database, which we generated for this exercise, we are going to create a table that can hold the information from the dataset that we have just explored:
USE [COVID-19] GO SET ANSI_NULLS ON GO SET QUOTED_IDENTIFIER ON GO CREATE TABLE [stg].[JHUConfirmedGlobal]( [ProvinceState] [varchar](100) NOT NULL, [CountryRegion] [varchar](100) NOT NULL, [Lat] [float] NULL, [Long] [float] NULL, [Date] [date] NOT NULL, [Value] [int] NULL, PRIMARY KEY CLUSTERED ( [CountryRegion] ASC, [ProvinceState] ASC, [Date] ASC )WITH (PAD_INDEX = OFF, STATISTICS_NORECOMPUTE = OFF, IGNORE_DUP_KEY = OFF, ALLOW_ROW_LOCKS = ON, ALLOW_PAGE_LOCKS = ON, OPTIMIZE_FOR_SEQUENTIAL_KEY = OFF) ON [PRIMARY] ) ON [PRIMARY] GO
I am creating the table in the stg (staging) schema, just in case later I would need to work a little more with the data in the database before being consumed by an application or report.
Load process
To take the information from the repository we are going to use Python and at some point, we will have to use Pandas also to unpivot some of the data that we receive as columns and pass it as rows. Pandas is a library that offers a large number of tools for working with structures and data manipulation within Python.
First, we are going to need the following libraries:
import os import requests import pyodbc as odbc import csv import pandas as pd
• os:
to use certain functionalities of the operating system to handle files.
• requests: to download the dataset.
• pyodbc: to connect to SQL Server.
• csv: to read the file separated by commas.
• pandas: to unpivot (convert columns into rows) the data for each date.
The rest of the code will look like this:
sql_conn = odbc.connect('DRIVER={ODBC Driver 17 for SQL Server}; SERVER=*******; DATABASE=COVID-19; Trusted_Connection=yes')
url='https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv'
response = requests.get(url)
with open(os.path.join("C:\Covid-19 Data Proyect\RawData", "time_series_covid19_confirmed_global.csv"), 'wb') as f:
f.write(response.content)
pd_data = pd.read_csv (r'C:\Covid-19 Data Proyect\RawData\time_series_covid19_confirmed_global.csv', encoding='utf-8')
pd_unpivoted_data = pd.melt(pd_data,id_vars=['Province/State','Country/Region','Lat','Long'], var_name='Date', value_name='Value')
pd_unpivoted_data.to_csv(r'C:\Covid-19 Data Proyect\RawData\time_series_covid19_confirmed_global_UNPIVOT.csv')
unpivoted_file = open('C:/Covid-19 Data Proyect/RawData/time_series_covid19_confirmed_global_UNPIVOT.csv','r', encoding='utf-8')
unpivoted_data = csv.reader(unpivoted_file, delimiter=',')
next(unpivoted_data)
sql_conn.execute('Truncate table stg.JHUConfirmedGlobal')
sql_conn.commit()
for row in unpivoted_data:
sql="""INSERT INTO stg.JHUConfirmedGlobal ([ProvinceState], [CountryRegion], Lat, Long, Date, Value)
values('"""+row[1].replace("'","´")+"""','"""+row[2].replace("'","´")+"""',"""+row[3]+""","""+row[4]+""",'"""+row[5]+"""',"""+row[6]+""")"""
print(sql)
sql_conn.execute(sql)
sql_conn.commit()
sql_conn.execute('exec stg.spNextProcess')
sql_conn.commit()
unpivoted_file.close()
sql_conn.close()
This is a basic example that we could use. At first, we downloaded the file and then we create another unpivoted file, we could delete the file after using it, but in this example, we suppose that we want to keep a history of the originals in the RawData folder since they weigh only a few KB and it is useful to be able to see what was it that we load in case of an error. In addition, the repository documentation clarifies that retroactive corrections can be made to the data, that is another good reason to save the originals for each day as it will allow us to make future comparisons. Knowing that the data can be corrected, the next step is to truncate the destination table, stg.JHUConfirmedGlobal, since we are going to do a full load instead of just loading the daily novelties. Then comes the insert loop. Finally, and if necessary, we can trigger any calculation or data movement that is necessary within the database. In the example, this is done by the stored procedure stg.spNextProcess.
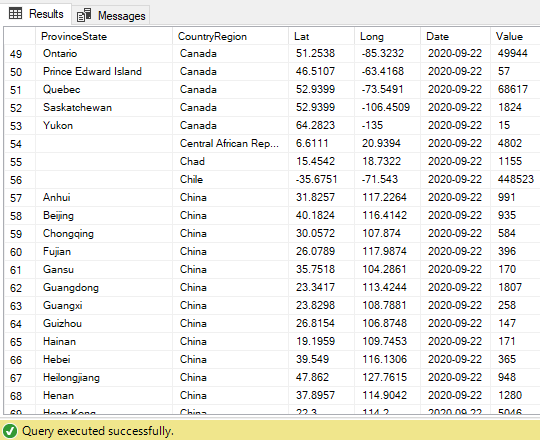
The end result will look like this. The Date and Value columns now holds the unpivoted rows data:
Scheduling the process execution
Suppose we want to execute the load every day, once a day, and suppose that this should happen early in the morning. It could be at 8 am, to make sure that by 9 am everyone is working with updated data. How do we achieve this? There are several ways to do it:
The first option that comes to our mind as SQL Server users is to create an Agent Job. For this, we are going to open the Management Studio (SSMS) and in the Object Explorer we are going to go to SQL Server Agent, Jobs, right-click and select New Job.
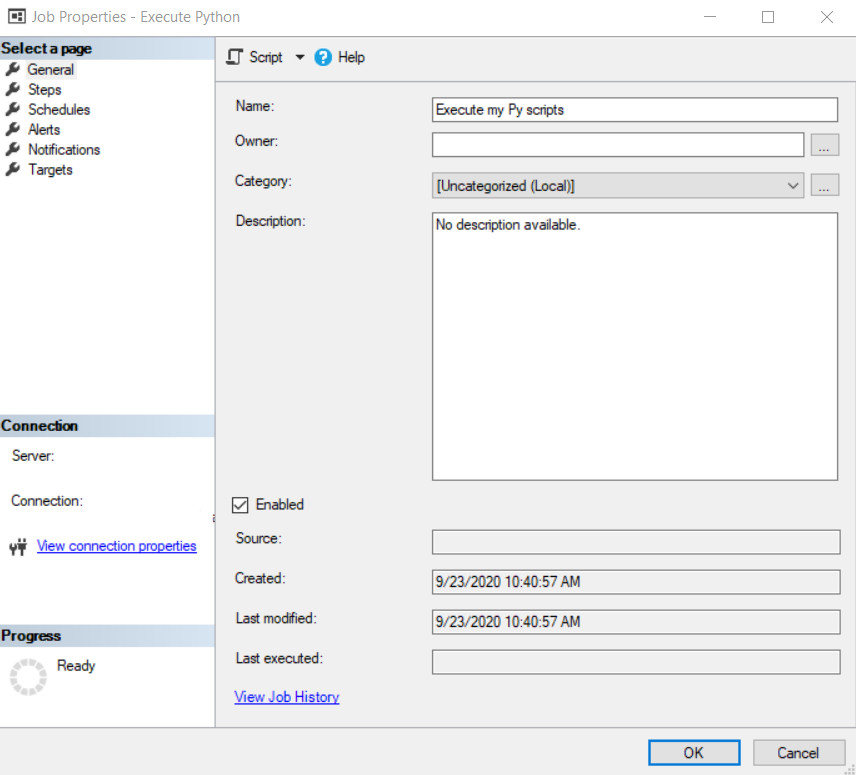
In the job we are going to create a step of type Operating system (CmdExec) and in Command we are going to write “C:\ProgramData\Python\Python38-32\Python.exe” “C:\Covid-19 Data Project\p01_load_confirmed_world.py” with the path and name of the script that you have created.
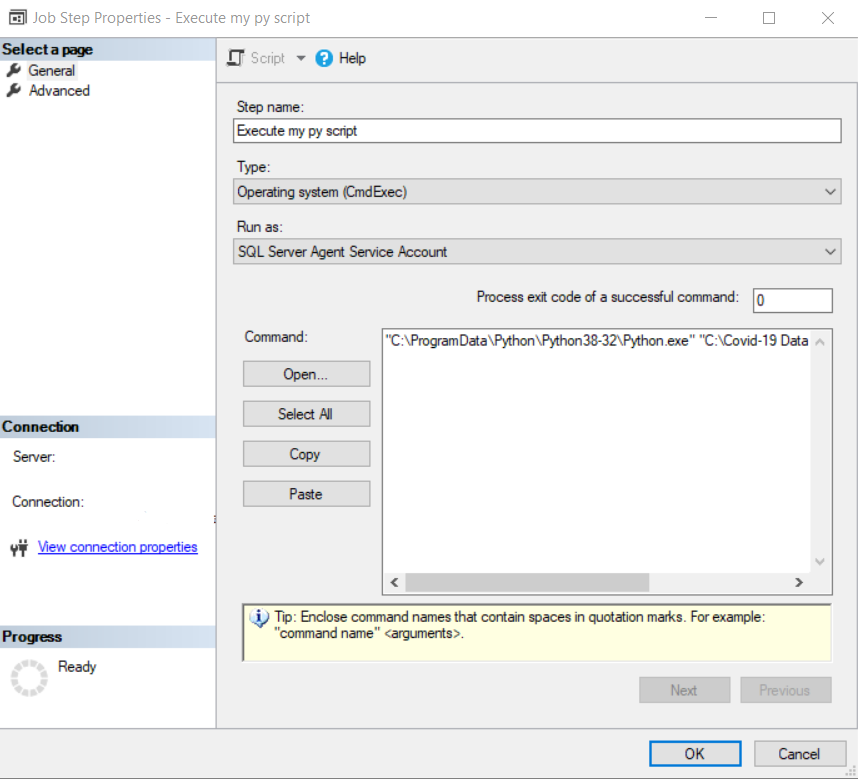
Finally, in the Schedules option, we are going to select the one we want to use, or we are going to create a new one for 8 am with the following options:
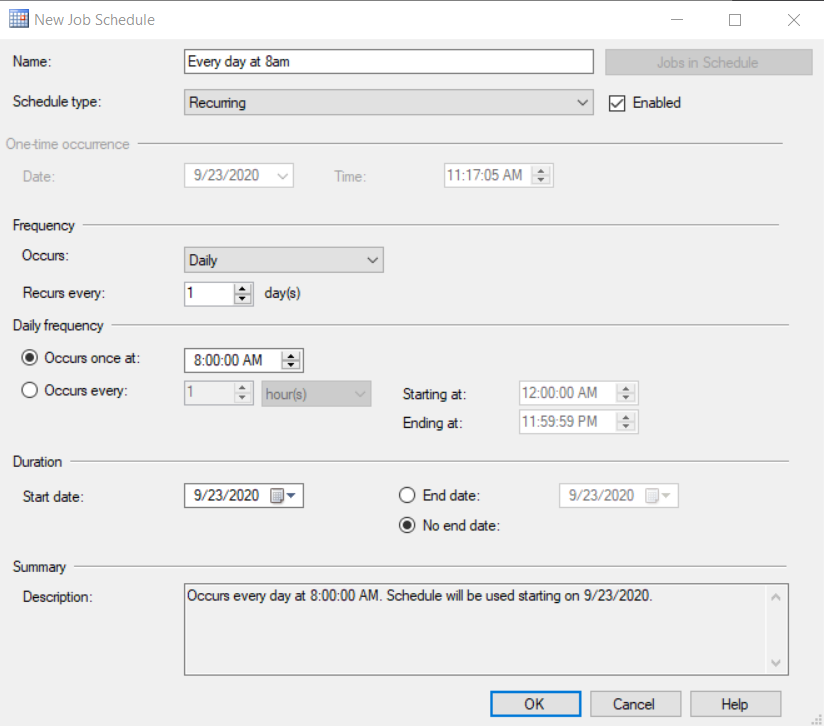
Of course, there are other scheduling tools if we do not want to use SQL Server jobs. A Python script can also be run from Windows Task Scheduler or from any other process scheduling tool such as ControlM.
With this last step, we have completed the development of the daily load. From now on, we only have to determine how we are going to use the information obtained so that it would be useful for end-users.
